
Ultra-large chemical libraries for the discovery of high-affinity peptide binders
Ultra-large chemical libraries for the discovery of high-affinity peptide binders
Nature Communications volume 11, Article number: 3183 (2020)
Anthony J. Quartararo, Zachary P. Gates, Bente A. Somsen, Nina Hartrampf, Xiyun Ye, Arisa Shimada, Yasuhiro Kajihara, Christian Ottmann & Bradley L. Pentelute
Abstract
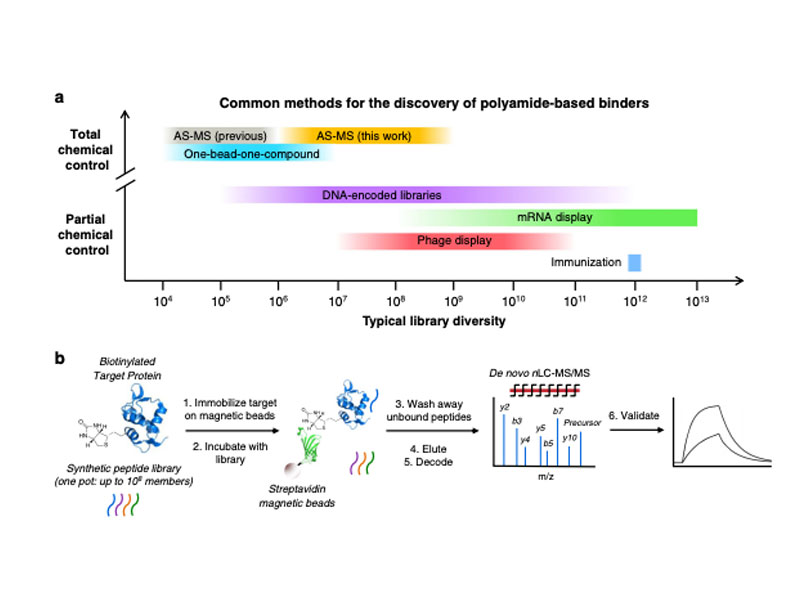
High-diversity genetically-encoded combinatorial libraries (108−1013 members) are a rich source of peptide-based binding molecules, identified by affinity selection. Synthetic libraries can access broader chemical space, but typically examine only ~ 106 compounds by screening. Here we show that in-solution affinity selection can be interfaced with nano-liquid chromatography-tandem mass spectrometry peptide sequencing to identify binders from fully randomized synthetic libraries of 108 members—a 100-fold gain in diversity over standard practice. To validate this approach, we show that binders to a monoclonal antibody are identified in proportion to library diversity, as diversity is increased from 106–108. These results are then applied to the discovery of p53-like binders to MDM2, and to a family of 3–19 nM-affinity, α/β-peptide-based binders to 14-3-3. An X-ray structure of one of these binders in complex with 14-3-3σ is determined, illustrating the role of β-amino acids in facilitating a key binding contact.



