
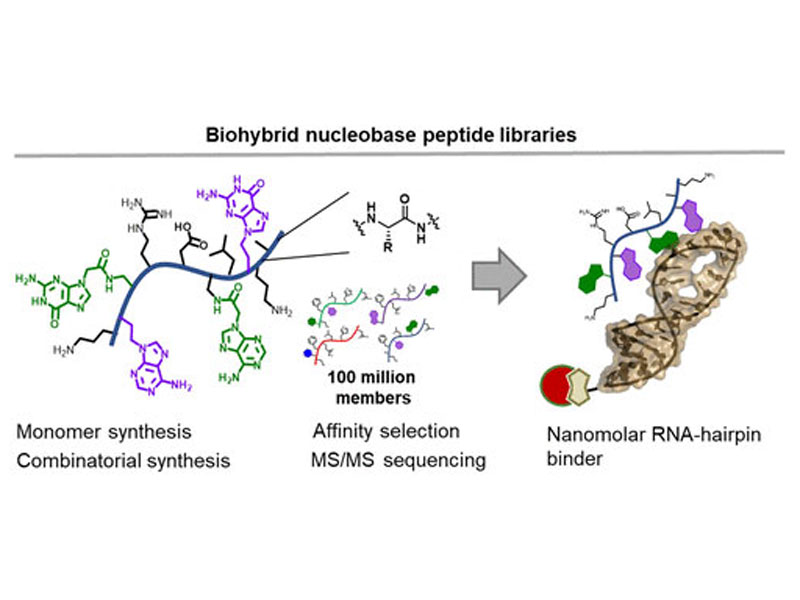
Discovery of Nucleic Acid Binding Molecules from Combinatorial Biohybrid Nucleobase Peptide Libraries
Discovery of Nucleic Acid Binding Molecules from Combinatorial Biohybrid Nucleobase Peptide Libraries
J. Am. Chem. Soc. 2020, 142, 46, 19642–19651
Publication Date:November 9, 2020
https://doi.org/10.1021/jacs.0c08964
Copyright © 2020 American Chemical Society
Sebastian Pomplun, Zachary P. Gates, Genwei Zhang, Anthony J. Quartararo, and Bradley L. Pentelute*
Abstract
Nature has three biopolymers: oligonucleotides, polypeptides, and oligosaccharides. Each biopolymer has independent functions, but when needed, they form mixed assemblies for higher-order purposes, as in the case of ribosomal protein synthesis. Rather than forming large complexes to coordinate the role of different biopolymers, we dovetail protein amino acids and nucleobases into a single low molecular weight precision polyamide polymer. We established efficient chemical synthesis and de novo sequencing procedures and prepared combinatorial libraries with up to 100 million biohybrid molecules. This biohybrid material has a higher bulk affinity to oligonucleotides than peptides composed exclusively of canonical amino acids. Using affinity selection mass spectrometry, we discovered variants with a high affinity for pre-microRNA hairpins. Our platform points toward the development of high throughput discovery of sequence defined polymers with designer properties, such as oligonucleotide binding.



